A*STAR NEWS
Sharing Data, Saving Lives: The Role of Bioinformatics During the Pandemic
Thinking out of the box to leverage technology and improve knowledge sharing globally
The COVID-19 pandemic is showing no signs of letting up, with rapid virus mutations disrupting lives and pushing hospitals in many parts of the world to near capacity. Key in the fight against the pandemic is the sharing of genomes of the virus, which helps with rapid response and the development of vaccines and diagnostic kits.
The Global Initiative on Sharing All Influenza Data (GISAID) platform, home to the largest SARS-CoV-2 sequence database in the world, has more than 9 million sequences today. The number is growing at a rate of about 1 million new sequences every five to six weeks.
A*STAR’s Bioinformatics Institute (BII), which specialises in computational biology and biomedical informatics has been working closely with GISAID to ensure that high-quality data and rapid analysis of the SARS-CoV-2 virus are shared globally in near real-time.
Executive Director Dr Sebastian Maurer-Stroh helms the research team at BII that has been working with GISAID for over a decade and round the clock for the COVID response since 2020. The multi-disciplinary team comprises domain experts such as Research Manager Mr Raphael Lee Tze Chuen, Post-doctoral Research Fellow Ms Shruti Vijay Khare and Senior Research Officer Ms Yani Xu.
 (From top) First row (from left): Louis DeFalco, Raphael Lee Tze Chuen, Joses Ho and Sandy Mak Tze Minn. Second row (from left): Chew Yi Hong, Roland Huber, Suma Tiruvayipati and Yani Xu. Third row (from left): Ashar Jamil Malik, Sebastian Maurer-Stroh, Fernanda L. Sirota and Shruti Vijay Khare. Fourth row: Meera Makheja
(From top) First row (from left): Louis DeFalco, Raphael Lee Tze Chuen, Joses Ho and Sandy Mak Tze Minn. Second row (from left): Chew Yi Hong, Roland Huber, Suma Tiruvayipati and Yani Xu. Third row (from left): Ashar Jamil Malik, Sebastian Maurer-Stroh, Fernanda L. Sirota and Shruti Vijay Khare. Fourth row: Meera Makheja
Global partnerships
During the early days of the pandemic between December 2019 and early January 2020, GISAID had just five genome sequences from three Chinese laboratories. Curating by the BII team was done manually. But by February 2020, manual curation was no longer an option as the virus had spread like wildfire. To ensure that genomes could be added quickly to the database, the Singapore team developed a software to automate the process.
In fact, from February to May 2020, A*STAR’s Genome Institute of Singapore (GIS) helped play a dynamic role in curating COVID-19 metadata and sequences, besides assisting in submitter and registered user requests.
As a volunteer researcher from GIS, Ms Suma Tiruvayipati served as the GISAID EpiCoV Singapore Curation Team Lead and laid the groundwork for COVID-19 sequence curation while aiding the BII team in the transition of GISAID EpiCoV's manual curation to its inception of automated curation.
Today, the joint BII/GIS team works together seamlessly to curate genomes, first checking for errors before adding them to the database in minutes. This quick turnaround is essential in the pandemic, enabling health officials and policy makers globally to make decisions that affect public health.
GISAID has ensured that every country has access to the software. In addition, the team submits regular reports that are widely used by researchers globally to monitor the pandemic and make timely tweaks to their local pandemic management strategies.
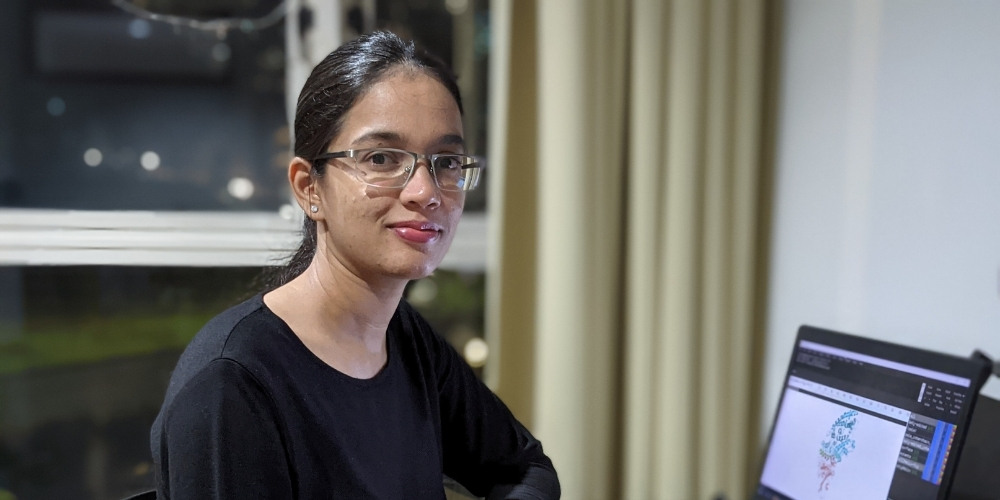 Ms Shruti Vijay Khare curates data for the GISAID database on a daily basis. Her frequent, real-time updates allow government officials, researchers and even the public to access information relating to the COVID-19 pandemic in a timely manner.
Ms Shruti Vijay Khare curates data for the GISAID database on a daily basis. Her frequent, real-time updates allow government officials, researchers and even the public to access information relating to the COVID-19 pandemic in a timely manner.
On a day-to-day basis, Ms Khare takes charge of curation at BII. Her work involves ensuring that good quality genome sequences are released to the users of GISAID. However, the increasing number of submissions in the last few months due to the spread of the Omicron strain has resulted in occasional bottlenecks in the curation process.
She observes that some users were uploading samples from the same patient as part of longitudinal studies.
To deal with the issue, she developed guidelines so users can indicate longitudinal sampling, and created a standard format which can be used to link different sample IDs from the same longitudinal sampling study.
Furthermore, she has been working closely with the software team to test new rules and provide feedback to continuously improve the curation process.
Creating new and better tools
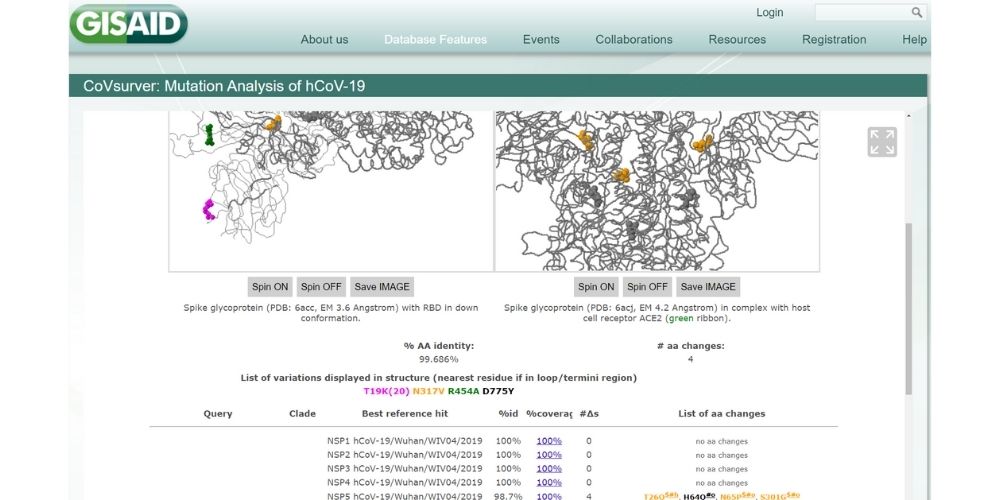 Innovated by BII, the CoVsurver is a key feature at the backend of the GISAID EpiCoV database, facilitating virus surveillance on the public and BII's end.
Innovated by BII, the CoVsurver is a key feature at the backend of the GISAID EpiCoV database, facilitating virus surveillance on the public and BII's end.
In January 2020, the team developed the CoVsurver tool that is used on the GISAID platform for sequence quality checks and annotations of every sequence submitted to the database, which helps to detect new virus strains. The tool eliminates manual analysis and increases accuracy of the data submitted.
The information is shared globally via GISAID, so public health officials can act in a timely way to control transmission and develop appropriate policies.
Using CoVsurver, the team helped Indian scientists in April 2021 to highlight the then new Delta variant and South African scientists in 2021 to analyse the first Omicron sequences. This includes singling out potential phenotypic effects of the variant.
Because of the speed at which the COVID-19 virus has been evolving, the Singapore team has been working with researchers globally and the National Supercomputing Centre Singapore (NSCC) to develop new computational algorithms to improve the quality and speed of some of the resources they provide to the global community.
On a regular basis, the team also provides data to international media organisations, as well as Covid sequence analysis reports to the World Health Organisation.
In late 2021, the team hosted a series of virtual bioinformatics workshops to show scientists from the ASEAN region, Africa, South America and Europe how to leverage the GISAID platform.
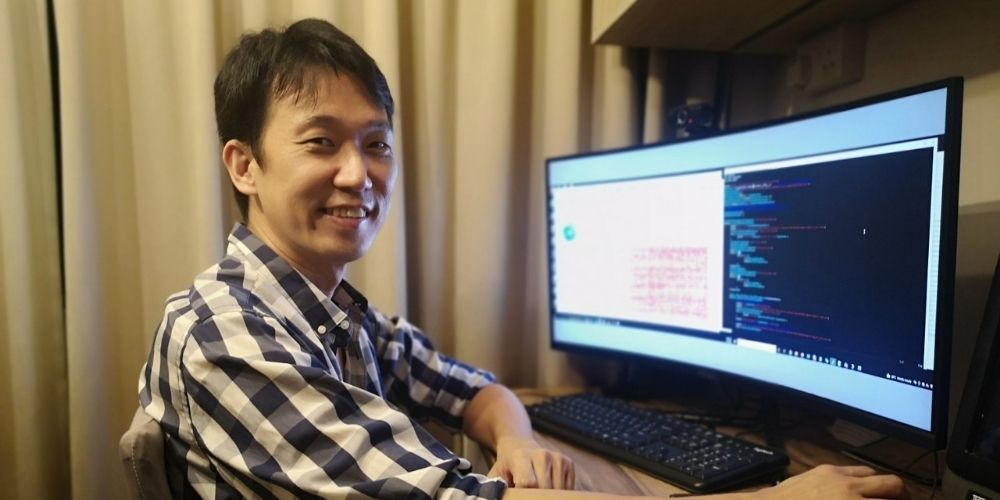 Mr Raphael Lee Tze Chuen is among the team that hosted a series of bioinformatics workshops to help educate some African nations and ASEAN countries on the GISAID platform.
Mr Raphael Lee Tze Chuen is among the team that hosted a series of bioinformatics workshops to help educate some African nations and ASEAN countries on the GISAID platform.
“Knowledge sharing is essential in the fight against COVID-19,” says Mr Lee. “At these sessions, we demonstrate to our peers how to prepare the genome sequences and metadata for submission to GISAID, how the curation process and CoVsurver work and how to use new tools like the dashboards we’ve created to detect new variants that spread quickly, and more.”
Thinking out of the box
As the pandemic continues, the BII team has been coming up with unique ways to analyse data, including genome sequences, even quicker.
It recently implemented a new automatic chart plotting feature in beta, which provides a snapshot of user search results, or epivocharts, from the GISAID EpiCoV database.
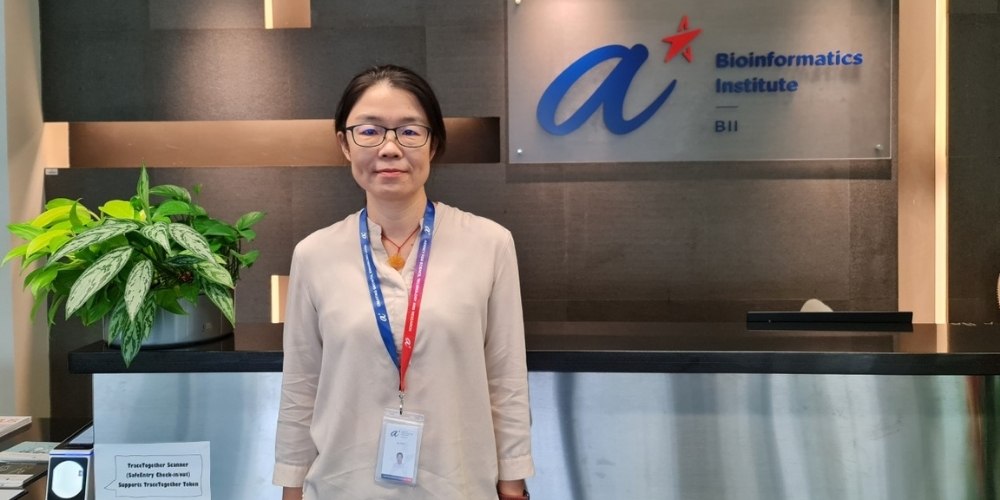 Following data curation, Ms Yani Xu creates charts, graphs and other useful graphics seen on the GISAID platform.
Following data curation, Ms Yani Xu creates charts, graphs and other useful graphics seen on the GISAID platform.
One of Ms Xu’s responsibilities is the development of epivocharts. She shares, “Since the GISAID EpiCoV database receives a lot of timely and valuable information about the virus from researchers around the world, we need to work around the clock and respond very quickly.”
For example, whenever a new variant of concern or variant of interest is announced by WHO, the team creates a new variant tracker page within the next few hours.
“This requires good teamwork from co-workers from different parts of the world,” she adds.
Staying the course
While the analysis of data and testing of genome sequences during the pandemic may be the bread and butter of the BII team, they are also focused on better understanding virus human-adaptation. The insights are valuable in preparing for pandemics of the future.
In addition, GISAID through BII provides regular reports to the Coalition for Epidemic Preparedness Innovations (CEPI) to monitor variants that could affect vaccines and advocate for equitable access to vaccines, lending a voice to less developed and developing countries that are struggling to control the pandemic.
More recently, the team has been helping scientists from different parts of the world in analysing virus samples that contain mutation signatures from different mixed lineages—helping these researchers to tell if they are due to co-infection of viruses from different lineages or truly new hybrid variants.
At the same time, the team is already closely tracking the spread of different Omicron sub-lineages and other potential variants of concerns, predicting if they might result in causing new waves of infection.
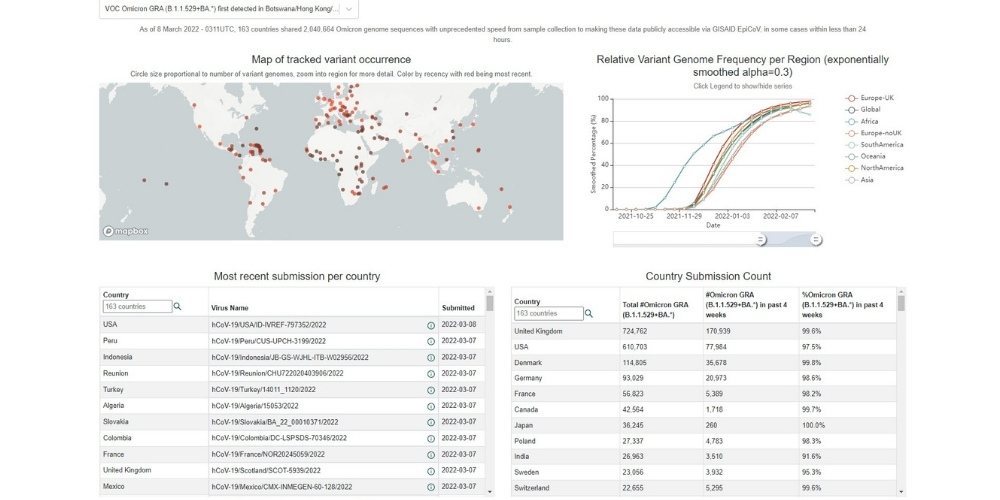 The BII team has developed a tracker to closely follow the spread of Omicron and other variants.
The BII team has developed a tracker to closely follow the spread of Omicron and other variants.
While much has been done, the war against the pandemic is far from over. The team continues to develop new and better ways of working, including building new tools and leveraging technology to automate processes to enable more efficient information sharing globally.
Was This Article Helpful ?
A*STAR celebrates International Women's Day

From groundbreaking discoveries to cutting-edge research, our researchers are empowering the next generation of female science, technology, engineering and mathematics (STEM) leaders.
Get inspired by our #WomeninSTEM
