A*STAR NEWS
PRESS RELEASES
Singapore scientists develop novel gene editor to correct disease-causing mutations into healthy versions
The CRISPR-based gene editor, CGBE, opens up treatment avenues for up to 40 per cent of genetic disorders caused by single-nucleotide mutations.
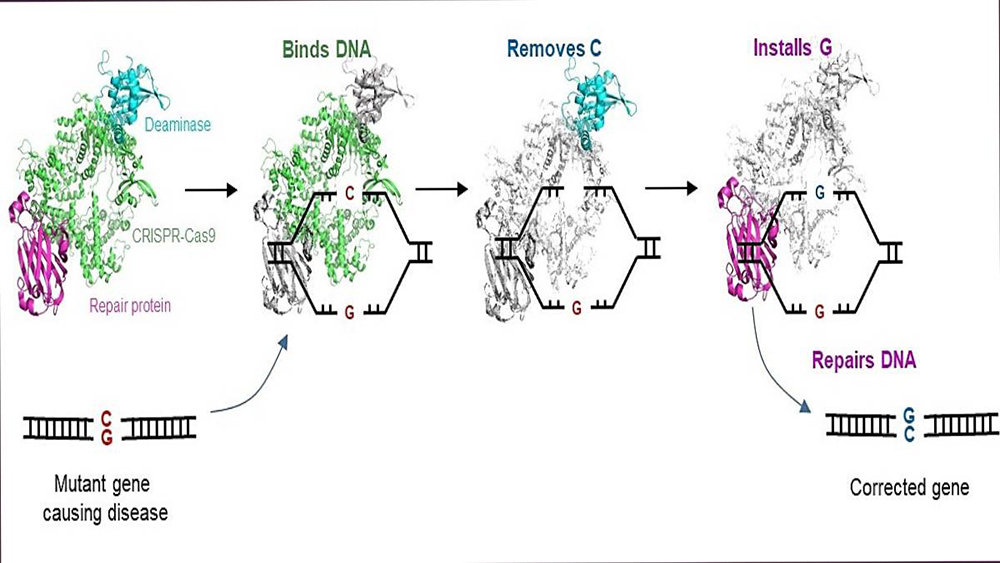
The C-to-G base editor (CGBE) converts C in genes to G. This invention corrects disease-causing mutations into healthy versions, enabling treatment for genetic diseases.
SINGAPORE – A team of researchers from Agency for Science, Technology and Research’s (A*STAR) Genome Institute of Singapore (GIS) has developed a CRISPR-based gene editor, C-to-G Base Editor (CGBE), to correct mutations that cause genetic disorders. Their research was published in Nature Communications on 2 March 2021.
One in seventeen people in the world suffers from some type of genetic disorder1 . Chances are, you or someone you know – a relative, friend, or colleague – is one of approximately 450 million people affected worldwide. Mutations responsible for these disorders can be caused by multiple mutagens – from sunlight to spontaneous errors in your cells. The most common mutation by far is the single-based substitution, in which a single-base in the DNA (such as G) is replaced by another base (such as C). Countless cystic fibrosis patients worldwide have C instead of G, leading to defective proteins that cause the genetic disease. In another case, replacing A with T in haemoglobin causes sickle cell anaemia.
To fix these substitutions, the team invented a CRISPR-based gene editor that precisely changes the defective C within the genome to the desired G. This C-to-G base editor (CGBE) invention opens up treatment options for approximately 40 per cent of the single-base substitutions that are associated with human diseases such as the aforementioned cystic fibrosis, cardiovascular diseases, musculoskeletal diseases, and neurological disorders.
The CGBE editor advances the widely adopted CRISPR-Cas9 technology to enable molecular surgery on the human genome. The CRISPR-Cas9 technology is routinely used to disrupt target genes, but it is inefficient when a precise change to particular sequences is desired. The CGBE editor resolves a key aspect of this challenge by enabling efficient and precise genetic changes. CGBE consists of three parts: 1) a modified CRISPR-Cas9, will pinpoint the mutant gene, and focus the entire editor on that gene; 2) a deaminase (an enzyme that removes the amino group from a compound) will then target the defective C, and mark it for replacement, and 3) finally, a protein will initiate cellular mechanisms to replace that defective C with a G. This enables a previously unachievable direct conversion from C to G, correcting the mutation and, consequently, treating the genetic disorder.
Dr Chew Wei Leong, Senior Research Scientist at GIS, said, “The CGBE gene editor is a ground-breaking invention that for the first time, directly converts C to G in genes, which potentially opens up treatment avenues for a substantial fraction of genetic disorders associated with single-nucleotide mutations.”
“The safety of patients is critical,” Dr Chew emphasised. “We are working to ensure our CGBE and CRISPR-Cas modalities are both effective and safe in disease models before we can further develop such modalities for the clinic.” For his scientific endeavours in gene editing therapy, he was one of the three young researchers that clinched the prestigious Young Scientist Award (YSA) 2020.
Prof Patrick Tan, Executive Director of GIS, said, “Novel editors such as CGBE are expanding the growing suite of precise genome-editing tools that include cytidine base editors (CBEs) 2, 3, adenine base editors (ABEs)4 , CGBEs 5, 6, 7, and prime editors . Together, they enable the precise and efficient engineering of DNA for research, biological interrogation, and disease correction, thereby ushering a new age of genetic medicine.”
– END –
About A*STAR’s Genome Institute of Singapore (GIS)
The Genome Institute of Singapore (GIS) is an institute of the Agency for Science, Technology and Research (A*STAR). It has a global vision that seeks to use genomic sciences to achieve extraordinary improvements in human health and public prosperity. Established in 2000 as a centre for genomic discovery, the GIS pursues the integration of technology, genetics and biology towards academic, economic and societal impact, with a mission to "read, reveal and write DNA for a better Singapore and world".
Key research areas at the GIS include Precision Medicine & Population Genomics, Genome Informatics, Spatial & Single Cell Systems, Epigenetic & Epitranscriptomic Regulation, Genome Architecture & Design, and Sequencing Platforms. The genomics infrastructure at the GIS is also utilised to train new scientific talent, to function as a bridge for academic and industrial research, and to explore scientific questions of high impact.
For more information about GIS, please visit www.a-star.edu.sg/gis.
About the Agency for Science, Technology and Research (A*STAR)
A*STAR is Singapore's lead public sector R&D agency. Through open innovation, we collaborate with our partners in both the public and private sectors to benefit the economy and society. As a Science and Technology Organisation, A*STAR bridges the gap between academia and industry. Our research creates economic growth and jobs for Singapore, and enhances lives by improving societal outcomes in healthcare, urban living, and sustainability. A*STAR plays a key role in nurturing scientific talent and leaders for the wider research community and industry. A*STAR’s R&D activities span biomedical sciences to physical sciences and engineering, with research entities primarily located in Biopolis and Fusionopolis.
ANNEX A – NOTES TO EDITOR
The research findings described in this media release can be found in the scientific journal Nature Communications, under the title, “Programmable C:G to G:C genome editing with CRISPR-Cas9-directed base excision repair proteins” by : Liwei Chen1, Jung Eun Park1, Peter Paa1, Priscilla D. Rajakumar1,3, Hong-Ting Prekop1, Yi Ting Chew1, Swathi N. Manivannan1, Wei Leong Chew1,*:
1. Genome Institute of Singapore, Agency for Science, Technology and Research, Singapore 138672, Singapore.
* Correspondence to: chewwl@gis.a-star.edu.sg
1 Jackson, M., Marks, L., May, G. H. W., Wilson, J. B. The genetic basis of disease. Essays Biochem 62, 643-723 (2018)
2 Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A., Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420-424 (2016).
3 Nishida, K. et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 353, aaf8729 (2016).
4 Gaudelli, N. M. et al. Programmable base editing of A*T to G*C in genomic DNA without DNA cleavage. Nature 551, 464-471 (2017).
5 Kurt, I. C. et al. CRISPR C-to-G base editors for inducing targeted DNA transversions in human cells. Nat. Biotechnol. (2020). https://doi.org/10.1038/s41587-020-0609-x.
6 Zhao, D. et al. New base editors change C to A in bacteria and C to G in mammalian cells. Nat. Biotechnol. (2020). https://doi.org/10.1038/s41587-020-0592-2.
7 Liu, D. R., Koblan, L. W. Cytosine to Guanine Base Editor. World Intellectual Property Organization (2018).
8 Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149-157 (2019).
Was This Article Helpful?
A*STAR celebrates International Women's Day

From groundbreaking discoveries to cutting-edge research, our researchers are empowering the next generation of female science, technology, engineering and mathematics (STEM) leaders.
Get inspired by our #WomeninSTEM
