Amit SINGHAL
Amit_Singhal@a-star.edu.sgResearch Interest
Host-Pathogen interaction and disease pathogenesis; Biomarkers and Point-of care diagnostics; Animal models; T cell biology; Host-directed therapy; Immunometabolism (Metabolic regulation during inflammation / cancer / infection)
Biographical details
Education
- Ph.D., All India Institute of Medical Sciences (AIIMS), New Delhi, India (2007)M.Sc.
- Animal Biotechnology, National Dairy Research Institute, Karnal, India (2000)
- B.Sc. Biochemistry, Kurukshetra University, India (1997)
Postdoctoral Training
- Singapore Immunology Network (SIgN), A*STAR, Singapore (2010 - 2012)
- Novartis Institute for Tropical Disease (NITD), Singapore (2008 - 2010)
- Institute Pasteur Brussels, Belgium (2007-2008)
Research Appointments
- Principal Investigator, A* ID Labs, A*STAR, Singapore (2020 - present)
- Principal Investigator, SIgN, A*STAR, Singapore (2017 - present)
- Adjunct Assistant Prof, LKC School of Medicine, NTU, Singapore (2016 - present)
- Adjunct Faculty, THSTI, Faridabad, India, (2017 - present)
- Project Leader, SIgN, A*STAR, Singapore (2014 - 2017)
- Senior Research Scientist, SIgN, Singapore (2012 - 2014)
Research Projects
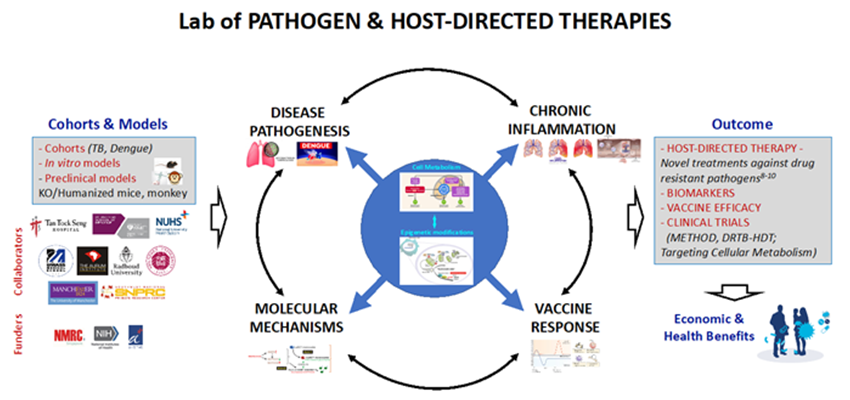
The AS Lab aims to dissect the complex dynamics of host-pathogen interactions following bacterial and viral infections during the innate and adaptive immune response phases. Main pathogens are Mycobacterium tuberculosis, Gram-negative bacteria and Dengue virus. We apply various omics approaches (at both population and single cell levels), to understand the molecular mechanisms utilized by pathogens for evading the host’s immuno-metabolic circuits. Since cellular metabolism plays Yin and Yang with epigenetic machinery, and epigenetic regulators help to establish innate and adaptive memory, we are particularly interested in chromatin-mediated transcriptional regulations. The information gained from these experiments is currently being exploited to design clinically relevant biomarkers and host-directed therapeutic (HDT) interventions (Fig 1).
TB Pathogenesis, Immunometabolism and Host-targeted Therapeutic Strategies
Mycobacterium tuberculosis (Mtb), the causative agent of tuberculosis (TB), is responsible for billions of infections worldwide. While treatable in 85% of cases, treatment is costly and relies on the patients’ adherence to a lengthy chemotherapy course, which results in non-compliance and eventually drug resistant TB. Drug resistance to first-line treatments is growing globally, and the requirement for new treatment options is becoming increasingly urgent.
Mtb targets immune cells for infection and replication. As such, understanding the pathways involved in Mtb infection/replication, as well as subsequent disease development and pathogen clearance, is central to the research conducted by the AS Lab. We utilize in vitro and in vivo models of infection along with ex vivo clinical samples to explore pathogen-induced perturbations in host cells from genomic to organismal level. This has led us to develop host-directed therapies (HDTs) which focuses on the activation of host factors subverted by the pathogens in order to improve disease pathogenesis and pathogen eradication. Such therapeutic approaches could potentially (i) augment the efficacy of antibiotic-based treatment, (ii) shorten treatment regimens for drug-susceptible and/or drug-resistant infections, (iii) reduce the immunopathology, and (iv) promote development of immunological memory that could protect against relapse and reinfection.
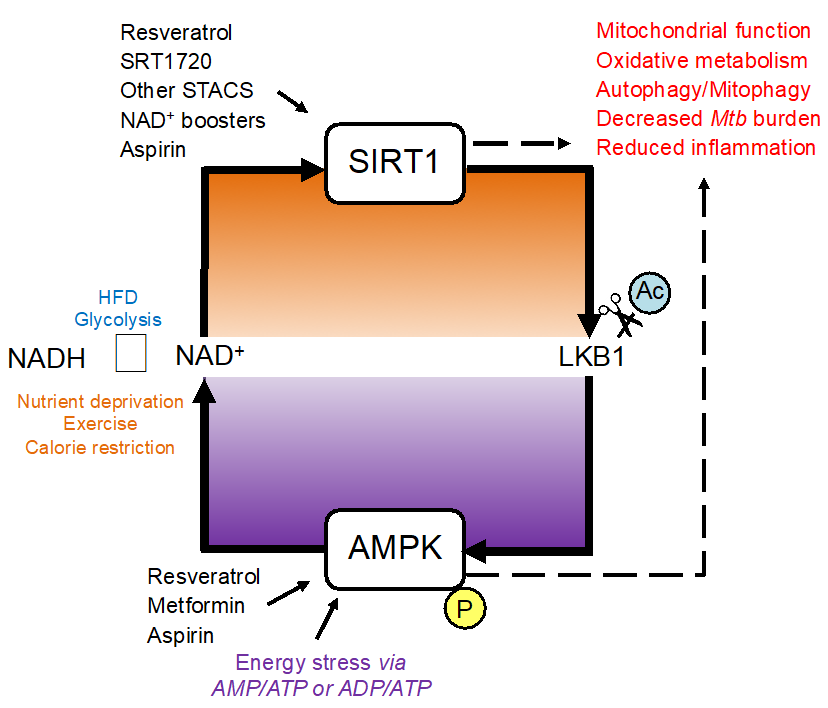
In this pursuit, we found potential of metformin (Singhal et al., 2014) and Sirtuin activators (Cheng et al., 2017) as TB-HDTs. These drugs target AMPK and SIRT1 respectively, which are important energy sensors and along with mTOR regulates host metabolic alterations and play an important role in immune homeostasis and responses against infections (Fig 2).
NAD+ been a crucial coenzyme in this crosstalk, we believe NAD+ metabolism can be intervened pharmacologically to enhance the host’s immunological fitness against pathogens.
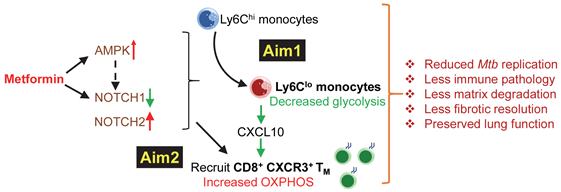
Metformin and immune cells: Our preliminary data and new findings (Bohme et al., 2020) indicate that metformin expands Ly6Clo non-classical (tissue repair) monocytes and memory CD8+ T cells (CD62L+CD44+, TM) in the lungs of uninfected and Mtb-infected mice. We hypothesize that this effect is due to metformin-mediated activation of AMPK and NOTCH signaling (Fig 3). Presence of Ly6Clo monocytes in the lung results in the recruitment of metabolically-primed CD8+CXCR3+ TM cells (Aim 2). Together this limit inflammation, matrix destruction and fibrosis and thereby preserve lung function in Mtb-infected hosts (Fig 3).
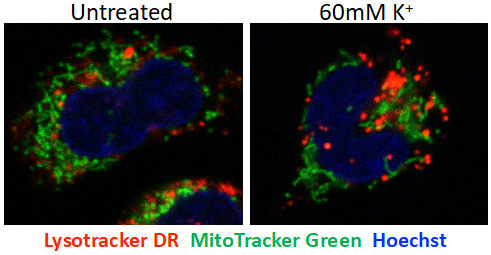
Potassium homeostasis and non-coding RNAs: Apart from AMPK and SIRT1 oriented investigations, other research in the lab focuses on understanding the role of long non-coding RNA (lincRNA), intracellular potassium fluctuations (Fig 4), and post-translational modifications such as phosphorylation, acetylation and ubiquitination during host-Mtb interaction. In these studies, approaches such as histone acetylome-wide association studies (HAWAS), single-cell ATAC / RNA sequencing, high throughput screening and machine-learning are being developed / utilized.
Monocytes and Dengue pathogenesis
Dengue virus (DENV) infects roughly 390 million people worldwide. During DENV infection, viremia does not overlap with the shock bleeding phase (critical phase), which indicate that other factors such as extensive inflammation in host contribute to the pathogenesis. Understanding the molecular mechanisms underlying the complex dengue-linked-inflammation will be helpful for improving therapeutic strategies for treating DENV infection. Since PBMCs from dengue-infected patients showed high percentage of infection in monocytes (49.5%), therefore, monocytes seem to be responsible for DENV pathogenesis. Furthermore, monocytes also promote tissue damage and vascular leakage, hallmark of severe dengue disease; and interact with DENV non-structural 1 (NS1), activating TLR4, thus support secretion of inflammatory cytokines. We aim to delineate which monocyte subset(s) contribute to the protective and pathogenic manifestations of DENV, and whether monocyte subset(s) are associated with clinical outcome. We utilize an unbiased single-cell transcriptomic approach (scRNAseq) and animal models to investigate these aims.
Nucleic acid therapies for targeting inflammation / infection
Systemic inflammatory diseases such as ARDS (Acute Respiratory Distress Syndrome), is an important cause of morbidity and mortality in intensive care units. Infections such as SARS-CoV-2 and Mycobacterium tuberculosis (Mtb) causes ARDS in patients with extensive pulmonary parenchymal involvement. Using an integrated analysis approach, based on Mtb modulated transcriptomic and epigenomic landscape in cellular models and patient derived material and in house developed CRISPR-Cas9 assisted homology directed recombination (CRISPR-Cas9-HDR), we have identified human-specific long non-coding RNAs (lncRNAs) as negative regulator of inflammatory responses. Using Locked nucleic acid (LNA) technology and human-specific in vitro models we aim to develop a lncRNA-targeting ASO (ltASO) to modulate acute and chronic inflammation / infection.
Role of NF-kB in acute myeloid leukemia (AML) and use of humanized mice
NF-kB family of transcription factors (TFs) help control key aspects of inflammation and are strongly linked to human pathological conditions including lymphomas and leukemia. Since, to date there is limited molecular data addressing the specific role of individual NF-kB TFs in human tumor, we designed a novel platform for gene knock-out (KO) using a gene synthesis independent paired gRNAs (pgRNA)-Cas9 lentivirus approach, to generate factor specific KOs of each of the five NF-kB TFs (p50, p52, p65, c-Rel and RelB) in the human monocytic/AML U937 cell line. RNA-sequencing in steady and inflammatory condition (LPS-stimulated) identified (i) state-specific homo- and heterogeneity in transcriptional regulation by individual members of the NF-kB family of TFs, and (ii) an inflammation independent previously undescribed unique biology of p65. We have developed humanized mice AML xenograft model which suggested that p65 regulated gene expression essentially captures AML-specific biological programs (metabolic plasticity and stemness), which appear to support leukemia development and progression.
Publications
Lab Members
| Postdocs (PhD) | Research Officers | PhD/Undergraduate Students |
|---|---|---|
| Wenning FANG | Andrea LEE | Saiji UEHARA |
| Yao CHEN | Mardiana Binte MARZUKI | Loukas PODAROPOULOS |
| Sheetal SAINI | LI Xinyi | Anabel CHANG |
| Sakshi AGARWAL | ||
Lab Photos
A*STAR celebrates International Women's Day

From groundbreaking discoveries to cutting-edge research, our researchers are empowering the next generation of female science, technology, engineering and mathematics (STEM) leaders.
Get inspired by our #WomeninSTEM
.png?sfvrsn=b11c2136_5)