RESEARCHERS CALL FOR STRONGER STANDARDS IN CANCER MICROBIOME STUDIES TO AVOID FALSE DISCOVERIES
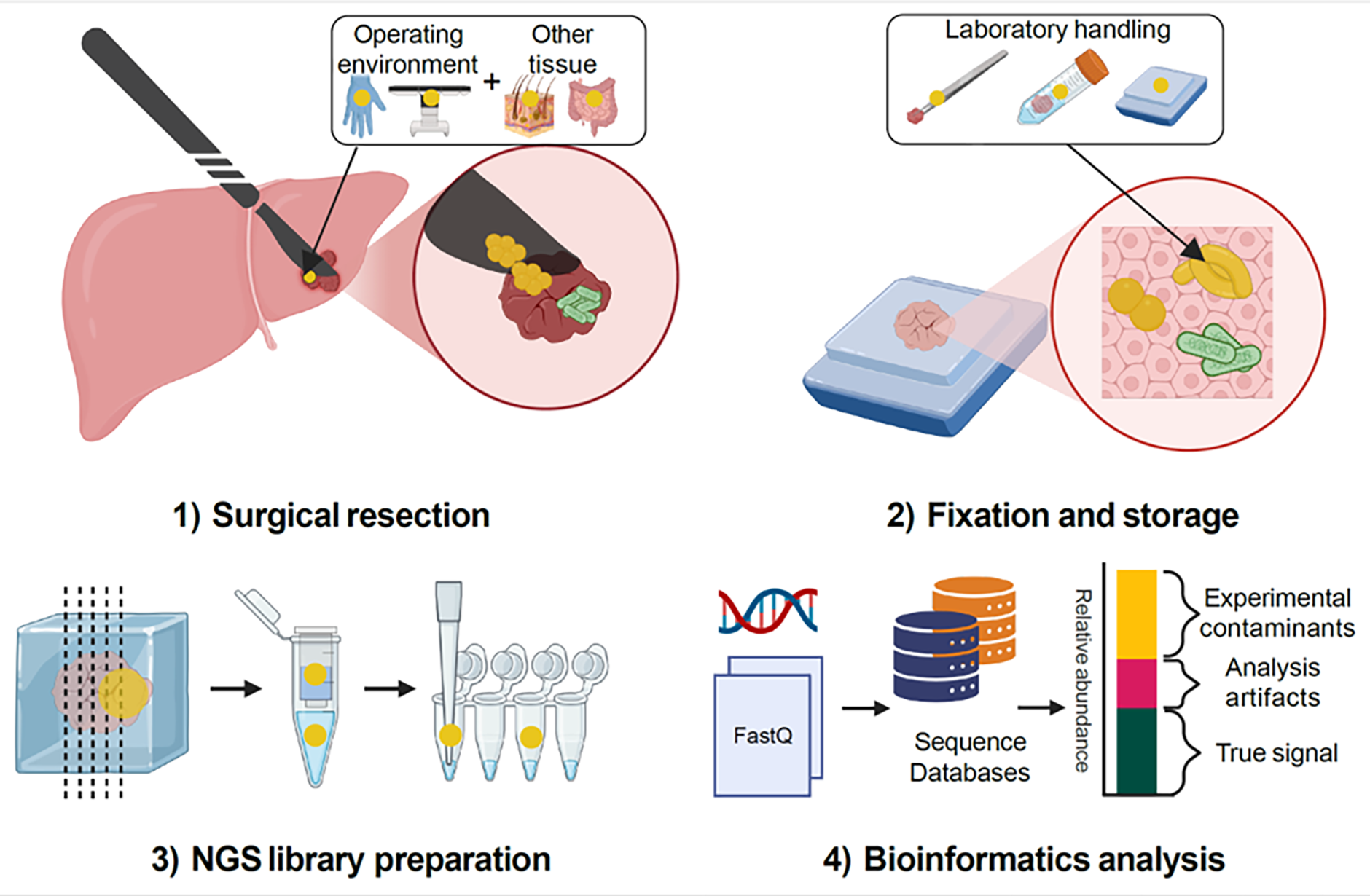
Possible sources of contamination in tumor tissues. Artefacts arising from environmental contaminants (yellow) and bioinformatics analysis (magenta) can confound interpretation of true microbial signatures (green) in tumor tissues, using liver cancer as an example. The impact of contamination is more significant at sites with lower microbial biomass such as breast, pancreas or liver. Created in BioRender: https://BioRender.com/v15scpn
Singapore, 13 March 2026 – A team of international scientists, led by researchers from A*STAR Genome Institute of Singapore (A*STAR GIS), has published a new perspective in Nature Cancer, calling for greater care and consistency in how scientists study microbes found in human tumours. Without proper controls, researchers may risk chasing false leads that could waste time, resources, and misdirect cancer research.
In recent years, scientists have reported finding communities of bacteria, fungi and viruses inside human tumours — sometimes in parts of the body where microbes are not usually expected, such as in the brain or in the kidney. These findings have sparked intense interest because some tumour-associated microbes may affect how cancers grow or respond to treatment.
But careful re-analysis of the data by various teams around the world have shown that many of these discoveries may not be as reliable as they seem. “Not all microbes reported in tumours are real residents,” said Assoc Prof Niranjan Nagarajan, Associate Director, AI & Compute at A*STAR GIS and lead investigator of this study. “Some could be the result of contamination or errors in data analysis. This could include microbial DNA accidentally introduced during sample handling or mistakes in analysis that wrongly matches human DNA to microbes.”
The researchers reviewed current practices in the fast-growing field of cancer microbiome research and identified several common pitfalls — from poor lab controls to issues with outdated databases. They argued that stronger standards and a checklist of recommended quality control steps are urgently needed.
To help the field move forward, the team has proposed a set of best practices that researchers can use when studying tumour-resident microbes. These include recommendations for preventing contamination, validating results using multiple methods, and being transparent about the limitations of the data.
This work is a result of an international collaboration between scientists from Singapore, the United States, the Netherlands, Denmark, and Australia — combining expertise in computational biology, genomics, microbiome science, and immunology.
Why This Matters
Microbes found in tumours could hold clues to why certain cancers grow faster, spread more easily, or respond differently to treatment. But false discoveries can distract from real progress and delay the development of better diagnostics and therapies.
“Imagine investigating a crime scene with contaminated evidence,” explained Dr Minghao Chia, GIS Fellow. “You might end up accusing the wrong suspect. That’s what can happen when researchers look for microbes in tumours without the right safeguards in place. By adopting more rigorous standards and confirming results with multiple approaches, scientists can avoid false discoveries, improve reproducibility, and focus research efforts on real connections between microbes and cancer rather than chasing misleading signals.”
Key Takeaways:
- Tumour-resident microbes may play important roles in cancer development and treatment.
- However, many published studies may be reporting microbes that are not truly present, due to contamination or analysis errors.
- The research team has proposed a clear set of guidelines to help improve the accuracy and reliability of future cancer microbiome studies.
The researchers hope their recommendations in the publication, Setting higher standards for reports of microbial species in human cancers | Nature Cancer, will raise the bar for research in this field and help uncover genuine biological links between microbes and cancer that can lead to real-world clinical advances.
###
About A*STAR Genome Institute of Singapore (A*STAR GIS)
The A*STAR Genome Institute of Singapore (GIS) is an institute of the Agency for Science, Technology and Research (A*STAR). It has a global vision that seeks to use genomic sciences to achieve extraordinary improvements in human health and public prosperity. Established in 2000 as a centre for genomic discovery, A*STAR GIS pursues the integration of technology, genetics, and biology towards academic, economic and societal impact, with a mission to harness genomic technologies for a better Singapore and world.
Key research areas at the GIS include RNA & DNA Technologies, Single Cell & Spatial Technologies), Precision Medicine, Population Genomics, and AI & Compute. The genomics infrastructure at the GIS is also utilised to train new scientific talent, to function as a bridge for academic and industrial research, and to explore scientific questions of high impact.
For more information about GIS, please visit www.a-star.edu.sg/gis.
Follow us on
LinkedIn | BlueSky | X | YouTube
About the Agency for Science, Technology and Research (A*STAR)
The Agency for Science, Technology and Research (A*STAR) is Singapore's lead public sector R&D agency. Through open innovation, we collaborate with our partners in both the public and private sectors to benefit the economy and society. As a Science and Technology Organisation, A*STAR bridges the gap between academia and industry. Our research creates economic growth and jobs for Singapore, and enhances lives by improving societal outcomes in healthcare, urban living, and sustainability. A*STAR plays a key role in nurturing scientific talent and leaders for the wider research community and industry. A*STAR’s R&D activities span biomedical sciences to physical sciences and engineering, with research entities primarily located in Biopolis and Fusionopolis. For ongoing news, visit www.a-star.edu.sg.
Follow us on
Facebook | LinkedIn | Instagram | YouTube | TikTok
.png?sfvrsn=2e525642_5)

