City-wide metagenomic surveillance of food centres reveals location-specific microbial signatures and enrichment of antibiotic resistance genes
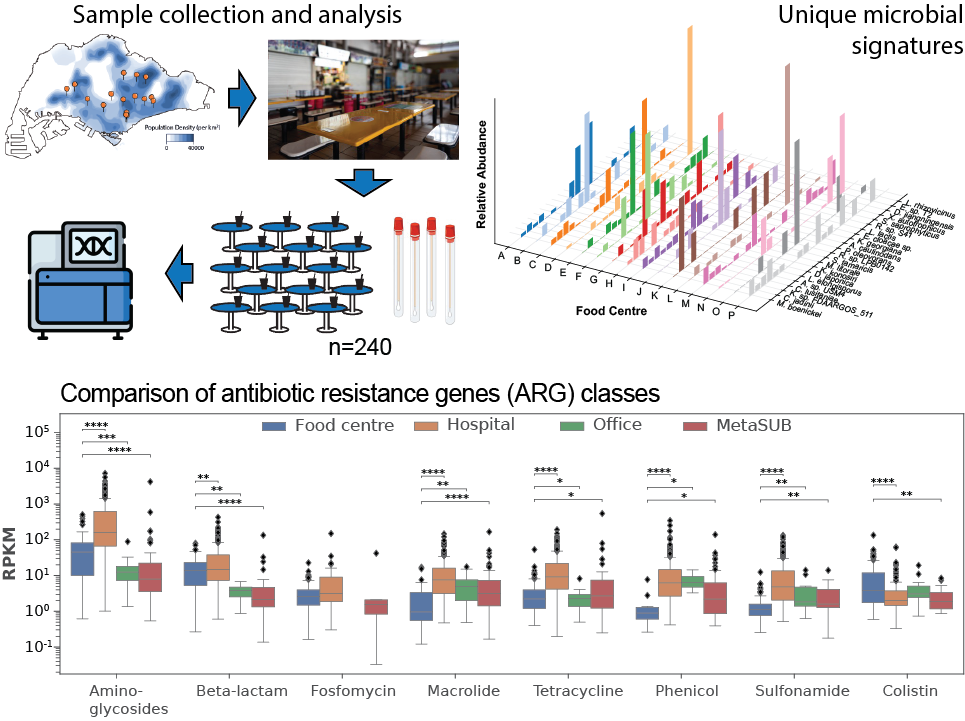
Top Left – Samples are collected from dining tables across 16 food centres in Singapore. Top right – Distinct microbial signatures are detected across food centres. Bottom - Antibiotic resistance gene (ARG) burden across antibiotic classes and locations are quantified as reads per kilobase per million mapped reads (RPKM).
6 September 2024 - Singapore's bustling food centres, famous for their vibrant hawker culture, have been discovered to host a diverse community of microbes that could have important implications for public health and food safety. A recent study by researchers at the Genome Institute of Singapore (GIS), in collaboration with the National Environment Agency (NEA) and Singapore Food Agency (SFA), marked the first large-scale effort to map the microbial life at these beloved dining spots.
The research team conducted an in-depth survey of 16 food centres across Singapore, using advanced DNA technology to uncover the invisible world of microbes that thrive there. These hawker centres, known for their wide variety of foods, also create an environment rich in nutrients that is favourable for a variety of microbes. The study found that microbes involved in food fermentation, like Bacillus and Exophiala, are commonly associated with a range of foods, suggesting they thrive on widely available nutrients. Another group of microbes, including Lactobacillus and Weissella, which are also involved in fermentation, were found to be linked to specific foods, suggesting their stricter nutritional requirement.
Beyond these findings, the study also revealed that each food centre has a unique microbial "fingerprint", a combination of specific microbes that sets it apart from others. Most microbial signatures, surprisingly defined by rare microbes rather than the more abundant ones, remained stable over time, even after three years. This consistency suggests that despite changes in the environment and human activity, the microbial communities in these food centres seem to persist. These unique microbial profiles could potentially be used to monitor changes in the microbial populations at food centres over time.
The research also highlighted the presence of certain microbial traits across food centres that are typically associated with different environments, such as hospitals. This discovery opens up new avenues for exploring the resilience and adaptability of microbial communities in diverse settings. By understanding how these communities exist and thrive, the research team can gain valuable insights that may inform future practices in maintaining healthy and stable microbial in food centres.
The authors emphasized the importance of continuing this research to further explore how these microbial communities could evolve over time and their potential impacts on the quality and safety in food centres. They also suggested that examining the effectiveness of current cleaning practices and developing new strategies could enhance the overall management of microbial populations in communal dining environments.
More details about their research paper at: https://www.medrxiv.org/content/10.1101/2024.07.28.24310840v1
A*STAR celebrates International Women's Day

From groundbreaking discoveries to cutting-edge research, our researchers are empowering the next generation of female science, technology, engineering and mathematics (STEM) leaders.
Get inspired by our #WomeninSTEM
.png?sfvrsn=2e525642_5)