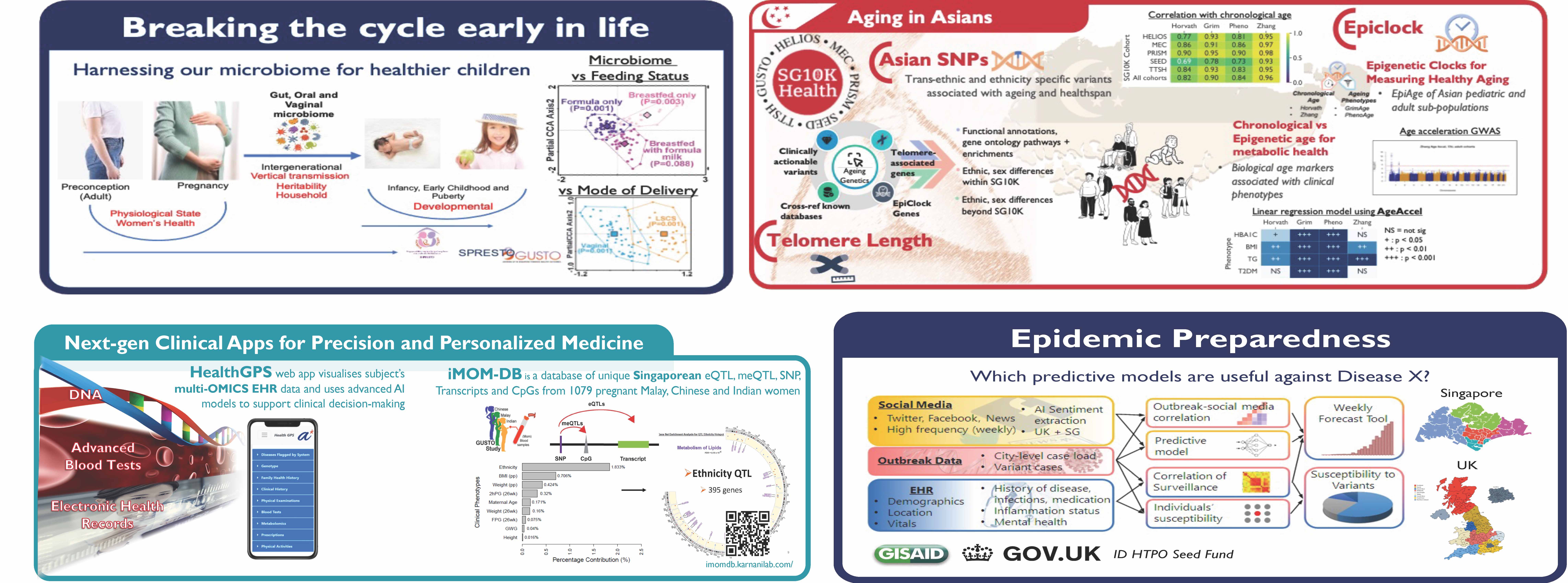
Clinical Data Engagement

Kanani Neerja
 | KARNANI Neerja Deputy Director (Clinical) / Senior Principal Scientist Email: neerja_karnani@a-star.edu.sg Research Group: Clinical Data Engagement |
Dr. Neerja Karnani is a molecular epidemiologist and clinical data scientist with 20 years of experience working with academia, industry, and national platforms. Dr. Karnani is a molecular epidemiologist and clinical data scientist with 20 years of experience working with academia, industry, and national platforms. She started her research career at the University of Virginia, USA, where she contributed to the first functional annotation draft of the human genome (ENCODE consortium). She moved to Singapore in 2013 to join Singapore Institute for Clinical Sciences (SICS), A*STAR. As the Systems Biology lead at SICS, she developed the multi-omics roadmap for Singapore’s National Birth (GUSTO) and Pre-conception (S-PRESTO) cohorts and identified biomarkers of metabolic and mental health adversities in expecting mothers and their offspring. These findings have been patented and licensed by eminent nutrition industries (Eg. Nestle and DSM) and are being translated into future interventions. She has also contributed to the research landscape and scientific vision of these cohorts by serving on their executive committees. Beyond the developmental and pregnancy cohorts, she is involved in Singapore National Precision Medicine (NPM) program’s SG10K-Health study and serves on its science and data access committees. She is also Co-chairing the ‘Aging in Asians’ study under the SG10K-Health initiative. To advance her interest in population health studies and to connect it with real-world data, she joined A*STAR’s Bioinformatics Institute (BII) in April 2021 as the Head of Clinical Data Engagement, and subsequently as the Deputy Director (clinical) and Biomedical Datahub Division Head. Under this new initiative, she is working closely with the Ministry of Health and public hospitals in Singapore to develop federated biodata hubs for chronic disease surveillance and epidemic preparedness. Her team is also developing nextgen mobile applications that can provide patient health journey and actionable intervention roadmaps for clinicians. She has authored over 100 publications covering various health domains of global concern and across life-course. Her contributions to the biomedical field and itsimpact on the public sector were highlighted in the GovInsider’s special international report on ‘Women in GovTech 2021’.)
Group Members
| BII Team | |
| Principal Scientist | GROZA Tudor |
| Senior Scientist | LIM Ives Yubin |
| Lead Research Officer | HUAN Jason |
| Research Officer | CHAN Penny |
| Data analysts under joint appointment @SICS | |
| Senior Scientist | GONG Min |
| Senior Scientist | XU Jia |
| Research Officer | TIN Felicia |
| Senior Research Officer | ONG Kelly |
A*STAR celebrates International Women's Day

From groundbreaking discoveries to cutting-edge research, our researchers are empowering the next generation of female science, technology, engineering and mathematics (STEM) leaders.
Get inspired by our #WomeninSTEM