Algorithms & Models of Protein Machinery
Su Tran-To Chinh
 |
SU Tran-To Chinh Research Scientist Email: chinhsutranto@bii.a-star.edu.sg Research Group: Algorithms & Models of Protein Machinery |
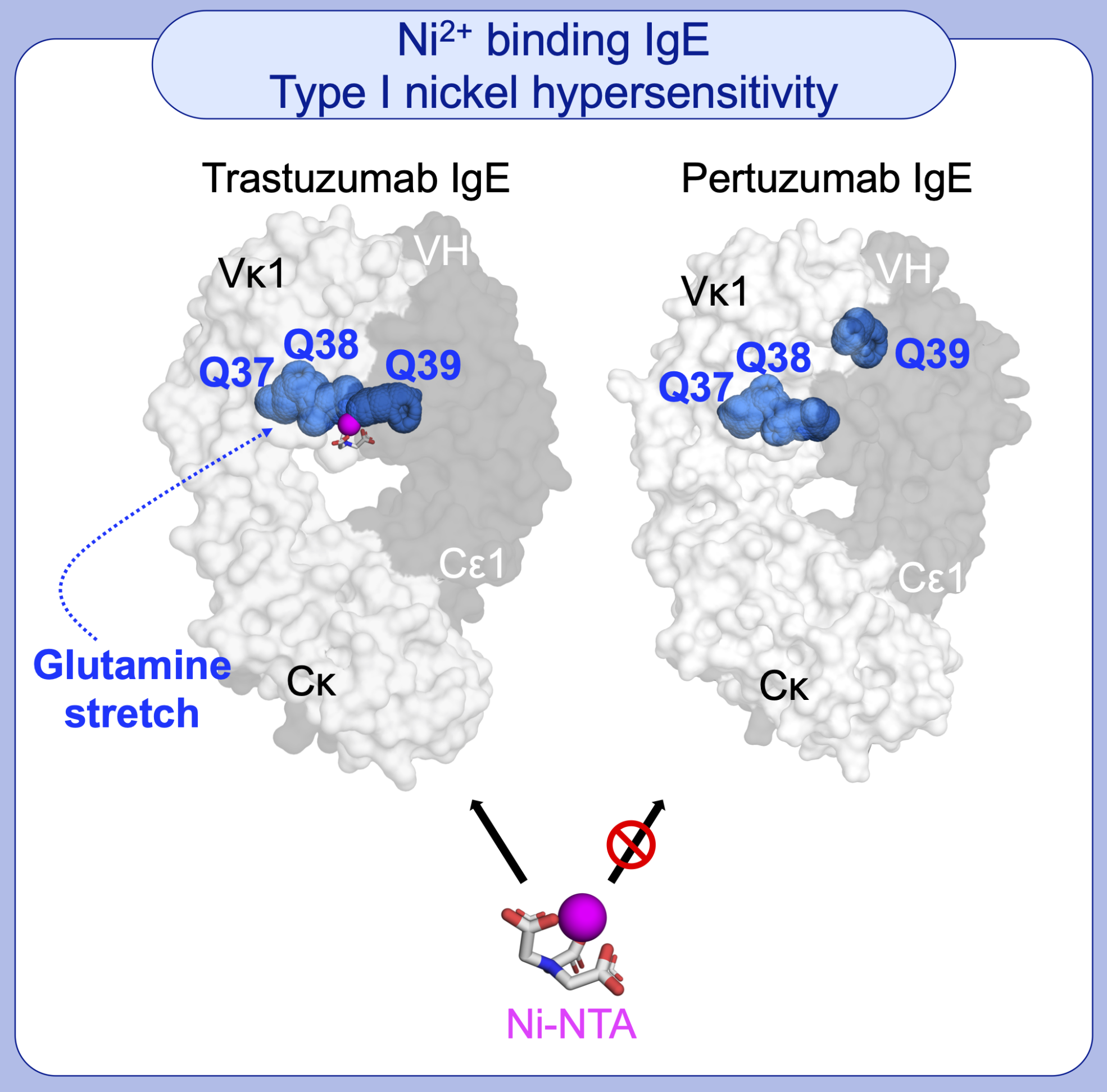
Chinh joined Bioinformatics Institute in 2014 after obtaining her PhD in Computational Biology from the School of Computer Science and Engineering, Nanyang Technological University. Chinh’s research areas focus on antibody and viral protein structural modeling and design, to which her work leverages on the fascinating interfaces of computational algorithms and structural models of protein complexes and interactions.
Chinh was awarded the BioMedical Research Council Young Investigator Grant (2015-2018) and the National Medical Research Council Open Fund Young Individual Research Grant (2021-2024) for research on structural modeling of proteins and their complexes to guide therapeutic interventions.
Research Interests
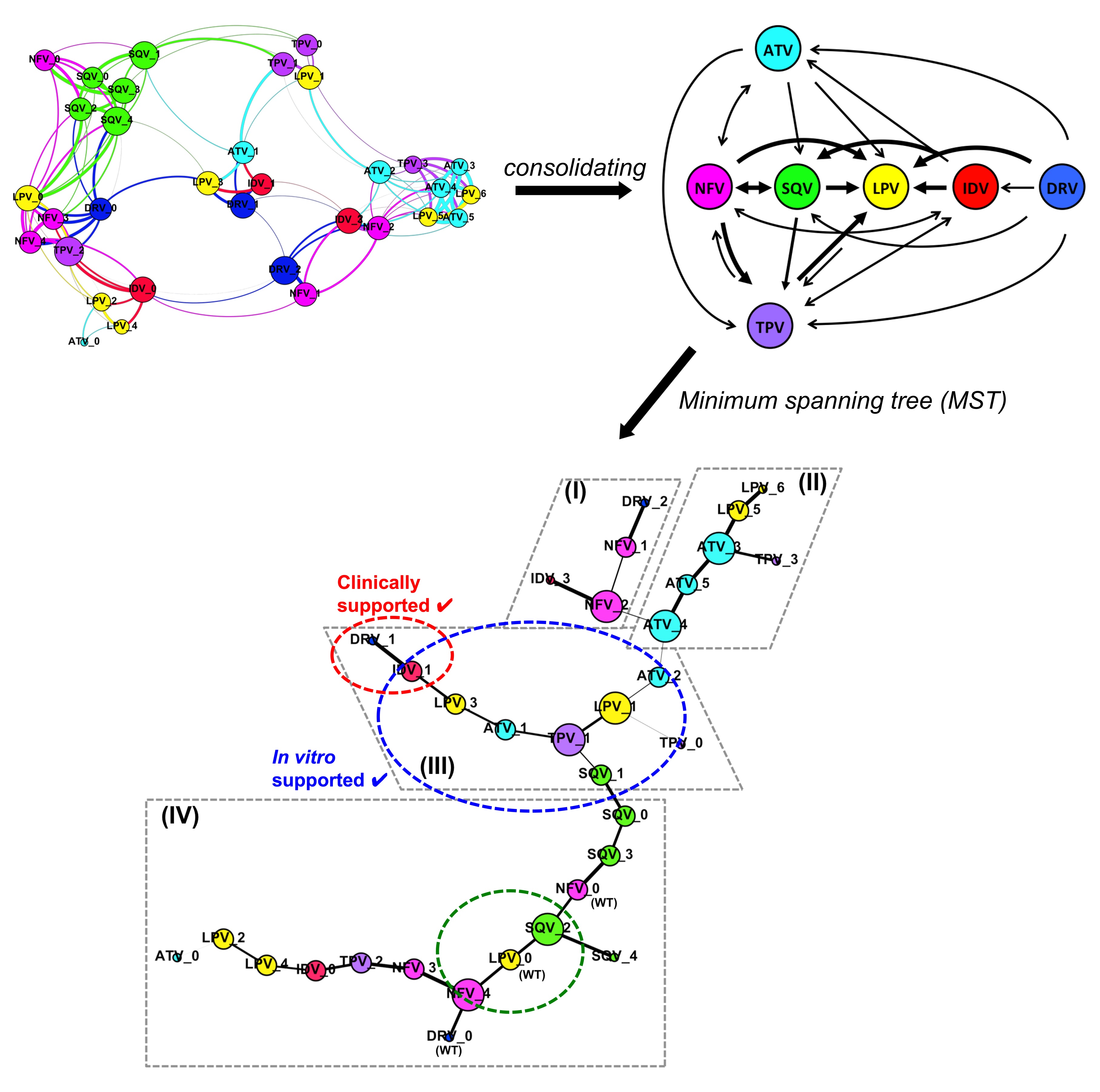
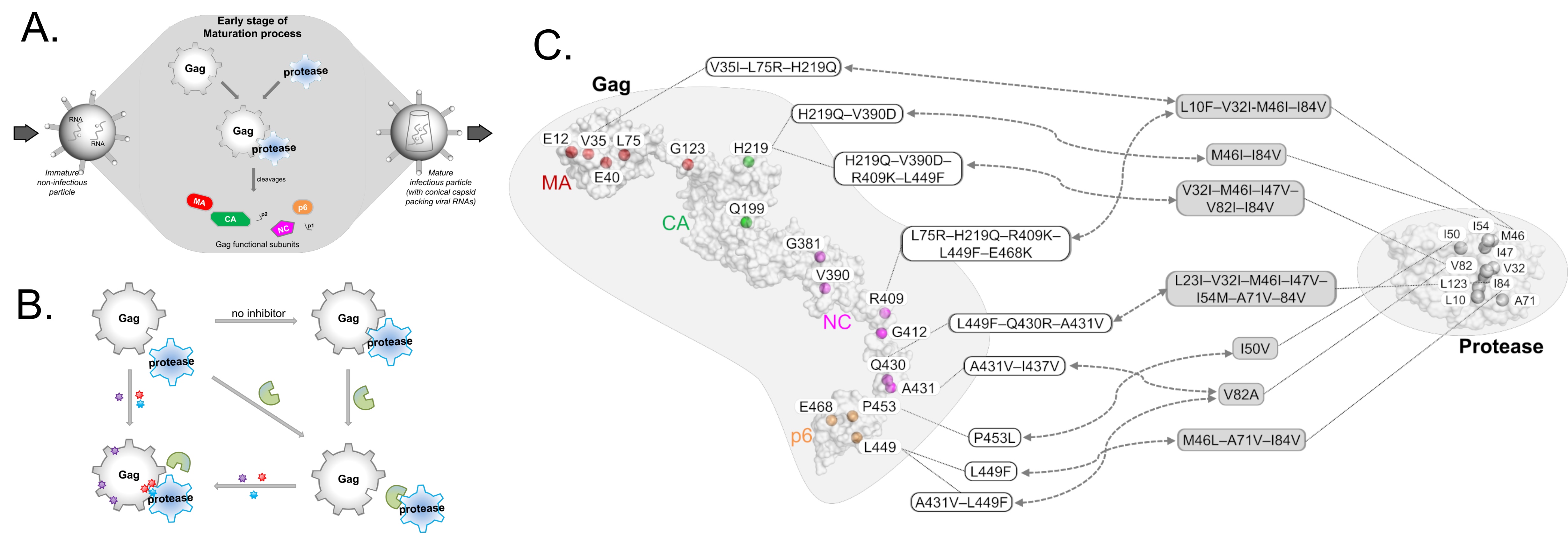
The group seeks to understand underlying mechanisms of how proteins of interest interact and how to modulate their interactions. We aim to look at protein interactions via the community concepts combined with protein structural dynamics at atomistic details.
For example, we apply network analysis on structural conformations of protein complexes to simulate their interaction networks or robotics-inspired algorithms (such as RRT sampling) to model possible conformational adaptation of protein domains. In addition, we aim to leverage on agent-based modelling on various conformational states of protein complexes and their networks to understand the interacting behaviors, striving for possible alternative insights useful for disease intervention design.
We also work with other computational and experimental collaborators for feedback on the modelled and devised systems.
A*STAR celebrates International Women's Day

From groundbreaking discoveries to cutting-edge research, our researchers are empowering the next generation of female science, technology, engineering and mathematics (STEM) leaders.
Get inspired by our #WomeninSTEM